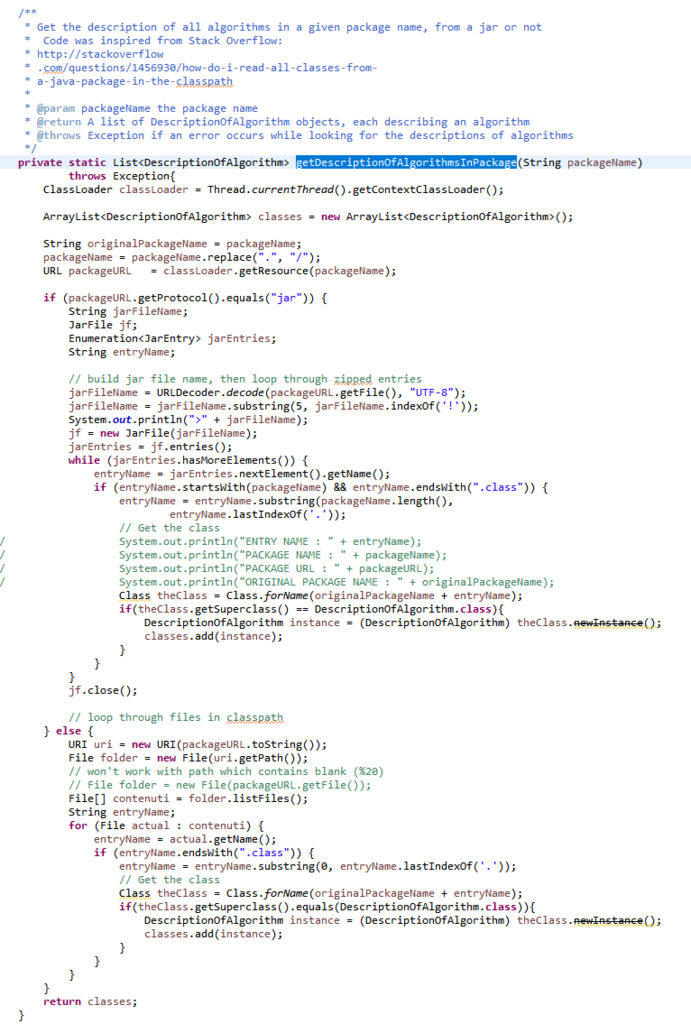
Today, I will talk about some efficiency problems when using the java.util.Hashmap data structure provided by Java. This is an important topics for programmers that aim at implementing efficient software that rely on maps.
Brief review about HashMap
First, let’s review why HashMap is important and what is special about this class. Hashmaps are important because they allow us to store and retrieve data efficiently and conveniently. More precisely, they store data in the form of key-value pairs, where each key is unique. The main feature of a HashMap is to allow efficient retrieval of a value in almost constant time O(1) if we know the key. Key-values pairs are stored in a hash table using a hash function. The most important operations of a HashMap are:
– put(key value) add a key-value pair to the map,
– get(key) retrieve a value from the map,
– remove(key) remove a key value pair from the map using its key
Moreover, some other operations are:
– contains(): to check if a key is in the map
Efficiency problems
Now, I will talk about some efficiency problems when using the java.util.HashMap implementation provided by Java.
(1) Handling of primitive types. First, many people will use a HashMap to store values from variable that are primitive types such as int, long, double, float, short. For example, one could write:
int x =0; int y =5; HashMap<Integer,Integer> map = new HashMap<Integer,Integer>(); map.put(x,y);
This will create a new key-value pair (0,5) in a HashMap for storing integers. This works but what is the problem with this? It is that Java will have to convert values of type int to Integer and we might have to convert again from Integer to int after retrieving a value from the map. This process called boxing and unboxing in Java is done automatically but it results in the unnecessary creation of more objects and thus a higher memory consumption.
Solution: A solution is to use an alternative implementation of HashMap that allows to directly use primitive types. There exists such implementations online. And this will be the case with some new data structures proposed in the next version of the SPMF software for example. In some cases, I have observed that using a custom map for primitive types can reduce memory use by up to 50%.
(2) Rehashing. A lot of interesting things can be learnt by looking at the source code of java.util.Hashmap. One thing that we can learn is that it is implemented as an array of buckets where collisions are handled using a linked-list for each bucket. And when the load factor (the number of elements stored in the map) exceed some threshold, the map is automatically resized to twice the size. This process of doubling the size of a HashMap allows to reduce the number of collision and thus maintain fast access to the key-values in the map. However, this process requires to do rehashing, that is to recompute the hash code of each key and reinsert all elements again in the map. Luckily, this is done quite efficiently as the objects representing the entries of the HashMap are reused when it is resized rather than creating new Entry objects. However, resizing a map still has a certain cost.
Solution: To reduce the cost of rehashing when using Java.util.HashMap, it is recommended to initialize the Hashmap to a proper size using a constructor such as:
/**
* Constructs an empty HashMap with the specified initial
* capacity and load factor.
*
* @param initialCapacity the initial capacity
* @param loadFactor the load factor
*/
public HashMap(int initialCapacity, float loadFactor);For example, if we expect an hashmap to contain 10,000 elements, we could initially set the size to a value greater than 10,000 and choose a load factor accordingly so as to reduce the cost of resizing().
(3) Handling collisions. As said above, the HashMap handles collisions using a linked-list for each bucket. And when there are too many collisions, HashMap can also switch to a tree structure for a bucket to reduce the time complexity of accessing entries. But generally, we should aim at reducing collisions.
Solution: To reduce collisions, one way is to choose an appropriate hash function that will assign keys to different buckets. To do this, we can define a custom hashCode() method. For example, if we want to put objects in a map that are instances of a class called Book as keys. Then, we could define an appropriate hashCode() function in the class Book to compute hash codes. Then, the HashMap will use that function to determine where the key,value pairs are stored. A good function will reduce collisions.
(4) Inefficiency on how the Hashmap is used. A big problem about HashMap is not about the structure itself but how it is used by the programmer to store and access key-value pairs. Let me show you a first example of that problem. Here is some code to store key-value pairs in a HashMap that are of type Integer,Integer:
HashMap<Integer,Integer> map = new HashMap<Integer,Integer>();
// Put some value 6 in the map for the key 5
map.put(5,6);
...
// Retrieve the value for key 5 and increase it by 1.
Integer value = map.get(5);
if(value == null){
map.put(5, 1);
}else{
map.put(5, value+1);
}
This code first puts a key-value pair (5,6) in the map. Then, later it retrieves the value for 5 and increase it by 1, and put it again in the map as (5,7).
What is wrong with this code? Well, it is that when map.get() is called, the hash code of 5 will be computed and the value will be searched in the HashMap. Then, when map.put() is called, the hash code of 5 will be computed again and it will be searched again in the HashMap. Doing this work twice is unecessary.
Solution: A better solution would be to have a method that would directly get the value in the map and increase it by the required value so as to avoid searching twice. In the early versions of Java ( < 1.8) I did not see an efficient way of doing this with the HashMap class provided by java.util. In the most recent Java versions, this can be achieved using the computeIfPresent() method.
But this can be also achieved if we use a custom Hashmap class. This is what is done for example, in the HashMap class that will be included in the next version of the SPMF software, with a method to do this directly:
void getAndIncreaseValueBy(int key, int valueToAdd); // Not in HashMap.
Here is a second example of inefficient programming with HashMap that I often see in the code of programmers that are not familiar with Java. The problem is to call several times the contains(…), get, and put methods. For example:
HashMap<Integer,Integer> map = new HashMap<Integer,Integer>();
...
if(map.contains(5) == false){
map.put(5,6);
}
What is the problem here? Well, by calling contains() and put(), the program will search twice for the value 5 in the Hashmap.
Solution: In this case, the solution is simple. We can just rewrite like this:
HashMap<Integer,Integer> map = new HashMap<Integer,Integer>(); ... map.put(5,6);
This code will do the same thing. If the key value pair (5,6) does not exist in the Map, it will be created, and if it does exist, then it will be replaced by (5,6). How do I know this? Because I have read the source code of HashMap to understand how it works 😉
A third example of inefficient programming with Hashmap is when someone needs to iterate over the key and values of a HashMap. For example, let’s say that we have a map called map. Then someone could fore example call map.entrySet().iterator() to obtain an iterator over all the entries of the Map. Or someone could call map.keySet().iterator() to obtain an iterator over all the keys of the map or map.valueSet().iterator() to obtain an iterator over all the values of the map. This can be OK if we are careful about how to use it. Some example of inefficient code is this:
HashMap<Integer,Integer> map = new HashMap<Integer,Integer>();
...
for(Integer key: map.keySet()){
Integer value = map.get(key);
...
System.out.println(" key-value: " + key + " - " + value);
}
What is the problem in this code? It is that it iterates on the keys and then call the get() method to obtain the corresponding value of each key. This is not necessary.
Solution.To avoid calling get, we should instead just do a loop using map.entrySet().iterator().
HashMap<Integer,Integer> map = new HashMap<Integer,Integer>();
for(Entry<Integer, Integer> entry: map.entrySet()){
System.out.println(" key-value: " + entry.getKey() + " - " + entry.getValue());
}
Using entrySet() we immediately obtain the corresponding value of each key without calling get().
Conclusion
java.util.HashMap is a very useful structure in Java but it should be used carefully. For this, I recommend to read the source code of HashMap to see how it is implemented (or at least to read the Javadoc) rather than to just use the methods that it provides without thinking. By reading the source code or documentation, we can understand how it works and write more efficient code.
Besides, the HashMap class is powerful but it also has some limitations. An example of limitation that I highlighted above is the inability to handle primitive values without auto-boxing (e.g. transforming int to Integer, or short to Short). To solve this problem, the solution is to use custom HashMap classes that handle primitive types.
Another issue with HashMap is that it provides a lot of features but sometimes all these features are not needed. Thus, sometimes a custom HashMap implementation that is lightweight would actually be more worthwhile when the goal is efficiency. For example, in the next version of the SPMF software that will be released soon, some custom HashMap classes will be provided for primitive types that are well-optimized to increase performance. I have observed that in some cases, it can reduce memory usage by over 50% for some data mining algorithms that do some intensive use of maps. But there are also drawbacks to using custom classes for HashMaps instead of the default one provided with one. One of them is that it may make the code a little harder to understand.
==
Philippe Fournier-Viger is a full professor and the founder of the open-source data mining software SPMF, offering more than 250 data mining algorithms.